Lab 5: Protein domain discovery & DNA motifs
Goals:
To identify 1. protein domains, 2. Gene Ontology (GO) terms, and 3. localization patterns in cells and/or tissues.
**Build slides with this information for your final talk.
To identify 1. protein domains, 2. Gene Ontology (GO) terms, and 3. localization patterns in cells and/or tissues.
**Build slides with this information for your final talk.
- Use at least Interpro (PFAM is found here now), SMART for protein domains (also try PROSITE)
- Gene Ontology: GO is great for multiple genes. GO terms can be found at Interpro as well.
- Model organism resources: Alliance of Genome Resources
- Human Protein Atlas
Sample slides to work from
| ataxin-2domains.pptx |
| ataxin-2domains.key |
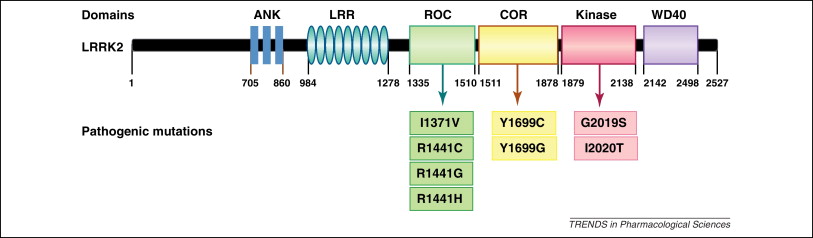
Example of a protein domain structure with mutations
Lab exercises
1)What Protein domains are present in your protein? (INTERPRO/SMART)
Build this in Powerpoint or Keynote.
Be sure to color & label each domain
KNOW what each domain does.
*see sample slides at top of page
Protein isoforms & mutation location:
Do you know if your protein has isoforms? If so, which domain(s) are missing in particular isoforms?
Do you know if your protein is alternatively spliced in your disease state?
Where do your mutations lie in the protein structure you found?
2. What is your Gene Ontology? Use Uniprot , Interpro or GO to find this information
Where does your protein or RNA localize (cellular component)?
What is it’s molecular function?
What biological process is your protein involved in?
3. Cellular component?
You can also find this on the Human Protein Atlas and via Pubmed
4. OPTIONAL: DNA motifs:
Using MEME
OPTIONAL but might be helpful depending on your project:
Gene Infinity: http://www.geneinfinity.org/index.html?dp=1
Are there any cis-Regulatory elements? Where is your promoter located?
Is your gene GC or AT rich? Any Repeats in your gene?
Does your gene have a Nuclear Localization Signal (NLS) or not?
Are there any Transcription factor binding sites?
Does your protein belong to a family or superfamily?
Is your protein Post-translationally modified?
--Example, is your protein phosphorylated and/or sumoylated? Find this under Protein Analysis on Gene Infinity Phosphorylation.
Build this in Powerpoint or Keynote.
Be sure to color & label each domain
KNOW what each domain does.
*see sample slides at top of page
Protein isoforms & mutation location:
Do you know if your protein has isoforms? If so, which domain(s) are missing in particular isoforms?
Do you know if your protein is alternatively spliced in your disease state?
Where do your mutations lie in the protein structure you found?
2. What is your Gene Ontology? Use Uniprot , Interpro or GO to find this information
Where does your protein or RNA localize (cellular component)?
What is it’s molecular function?
What biological process is your protein involved in?
3. Cellular component?
You can also find this on the Human Protein Atlas and via Pubmed
4. OPTIONAL: DNA motifs:
Using MEME
OPTIONAL but might be helpful depending on your project:
Gene Infinity: http://www.geneinfinity.org/index.html?dp=1
Are there any cis-Regulatory elements? Where is your promoter located?
Is your gene GC or AT rich? Any Repeats in your gene?
Does your gene have a Nuclear Localization Signal (NLS) or not?
Are there any Transcription factor binding sites?
Does your protein belong to a family or superfamily?
Is your protein Post-translationally modified?
--Example, is your protein phosphorylated and/or sumoylated? Find this under Protein Analysis on Gene Infinity Phosphorylation.