| gen564interactionlab2023.doc |
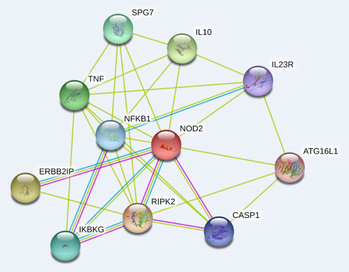
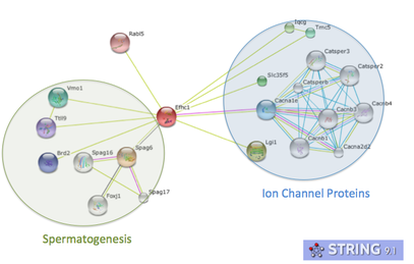
Lab 9: Building Protein Interaction Networks
Using your favorite protein of interest (or at least a well-known protein such as NOTCH or CDC2 for purposes of familiarizing yourself with the databases), find previously identified interacting proteins using a few databases listed below. If your protein of interest has a mouse, yeast, fly, worm, or zebrafish homolog check out the protein interaction data for it.
What interaction networks can help you understand: If you are still unsure what your protein does, you might be able to infer what your protein does by the proteins that it interacts with. Did you find any proteins that interact with your protein that you find interesting based on the biological function of this identified protein, etc.??? Do they share similar protein domains? Do any of the GO term biological functions suggest that this protein(s) may regulate your protein in some way or via other pathways that open new avenues of research? For example, if you have an ER protein and find that it interacts with a cell cycle regulator what can this tell you? (hint: the ER protein either is regulated during the cell cycle or regulates this cell cycle protein, etc.).
What interaction networks can help you understand: If you are still unsure what your protein does, you might be able to infer what your protein does by the proteins that it interacts with. Did you find any proteins that interact with your protein that you find interesting based on the biological function of this identified protein, etc.??? Do they share similar protein domains? Do any of the GO term biological functions suggest that this protein(s) may regulate your protein in some way or via other pathways that open new avenues of research? For example, if you have an ER protein and find that it interacts with a cell cycle regulator what can this tell you? (hint: the ER protein either is regulated during the cell cycle or regulates this cell cycle protein, etc.).
- Find your interaction networks for your Human protein and at least in your model organisms (yeast, mouse, fly, worm, zebrafish, Arabidopsis). Remember to use the organism specific name if you are looking for your homolog in that particular model organism or use FASTA sequences.
- Determine the Gene Ontology (cellular component and/or biological process) for each protein interactor. You can use STRING directly (Under Analysis Tab). Click on whatever GO term list you like.
*** Remember you will get different results for different model organisms—Yeast, fly, worm, fish, mouse, plant will have more data. So, try your homologs and see if you see anything different between what is known in humans vs all of your homologs. When you choose a certain model organisms in your aims, and use interaction networks pay close attention to similarities and differences. - Place this data on your website under your protein section, build a slide for your talk that shows your network for human and your model organism choice, and sort your protein interactions based on gene ontology (by placing a transparent color coded circle by Gene Ontology Groups on top of your sorted groups—which can be done in PowerPoint or Keynote then exported as an Image file (jpg or PNG) and upload to your website.___________________________________________________________________________________________________________
- Image saving: To save interaction networks, find the Tables/Exports below your network and save as .png file likely a low resolution one will work fine on your website. Note the edge colors, as this will tell you how the data was found.
- Save as Excel file to easily identify your GO terms for all interactors:
Under Tables/exports (see image above). Choose ….as simple tabular text output. Open in Excel, now copy and paste your Accession codes into GO, Uniprot or Panther to identify all biological processes for all your interactors quickly. - Gene Ontology Terms can be found in String: Under Analysis you can easily download your 3 GO terms associated with the interactors you find. Go to the bottom of the page and you can export your data for use in excel.
- Other Protein Interaction Database Websites
BioGRID: http://www.thebiogrid.org
I use both String and BioGrid often. Check out the Network tab (like visual in String) and different ways to visualize the interaction network here.
IntAct: http://www.ebi.ac.uk/intact/index.jsp
This page is not as visually appealing but has similar data. Check the graph tab to visualized the interactions.
GO analysis
PANTHER is easier for batch searching a list of Protein Ontologies: http://www.pantherdb.org